Cell Stem Cell | Teams of Yawei Gao and Jun Liu Reveal Mechanism of m⁶A-mediated LTR Silencing and Totipotency Restriction via L1PA


Cell Stem Cell | Teams of Yawei Gao and Jun Liu at SRILS Co-publish Research Revealing m⁶A-mediated LTR Silencing via L1PA RNA
In early mammalian embryonic development, the first "self-voice" of life originates from Zygotic Genome Activation (ZGA)—the earliest watershed of cell fate. With the onset of ZGA, the embryo shifts from maternal transcriptional control to autonomous transcription. Cells transition from a totipotent state to a naïve pluripotent state and then to a primed pluripotent state, a transformation that not only defines the developmental trajectory but also provides a blueprint for in vitro stem cell models.
During this process, the activation and silencing of repetitive sequences—particularly Long Terminal Repeats (LTRs)—constitute the hidden rhythm of the developmental program. These elements serve as both markers of totipotency and targets for epigenetic regulation. While previous research by Prof. Shaorong Gao's team emphasized the role of H3K9me3 in LTR silencing, the role of RNA-level modifications, especially m⁶A (N6-methyladenosine), in human ESCs remained unclear.
On October 29, 2025, teams led by Prof. Shaorong Gao, Prof. Yawei Gao, Prof. Yixuan Wang, and Prof. Jun Liu published a study in Cell Stem Cell titled "N6-methyladenosine on L1PA Governs the Trans-silencing of LTRs and Restrains totipotency in Naïve Human Embryonic Stem Cells."

By using METTL3 inhibitors to reduce m⁶A levels in naïve human ESCs, the researchers observed a slowdown in cell proliferation and a significant upregulation of 8-cell stage (8C) marker genes. METTL3 inhibition blocked the differentiation from naïve to primed states, instead promoting a transition toward an 8C-like totipotent state. Transcriptomic and single-cell analysis confirmed that m⁶A inhibition triggers widespread totipotent transcriptional activation, particularly within ERV1 and ERVL-MaLR subfamilies.
Further nascent RNA-seq and ATAC-seq revealed that the transcriptional activity and chromatin accessibility of 8C genes, eRNAs, and LTRs increased significantly. The study established L1PA as a key m⁶A target. Using the FTO-dCas13b system to precisely erase m⁶A on L1PA replicated the transcriptomic features of global METTL3 inhibition.
ChIRP-seq results showed that L1PA RNA is enriched in enhancer and 8C LTR regions, directly binding to LTRs and recruiting epigenetic factors. The team proposed that m⁶A acts as a "selector" for histone modification regulators: m⁶A-modified L1PA recruits KAP1 to maintain H3K9me3 silencing, while the loss of m⁶A promotes EP300 binding and H3K27ac accumulation.
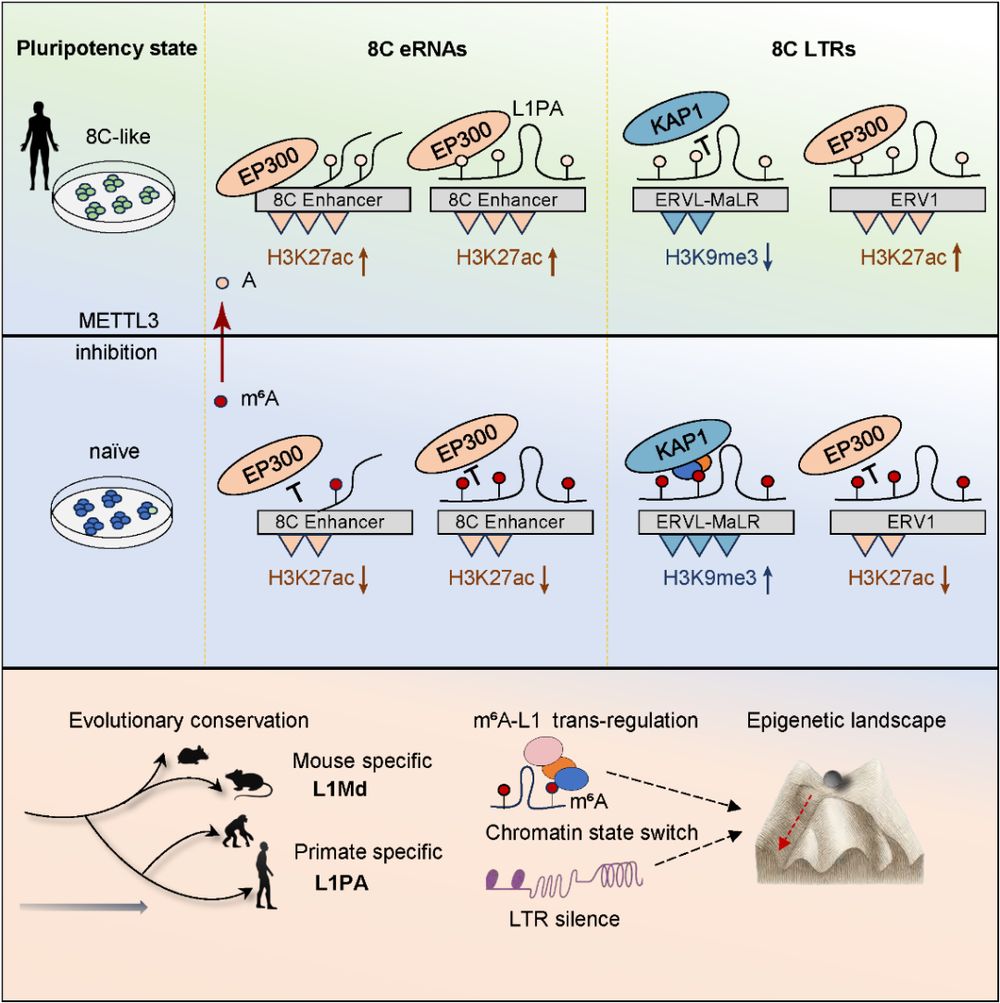
Overall, this study identifies a hierarchical regulatory network—the m⁶A–L1PA–LTR axis—as a critical bridge between RNA modification and epigenetic remodeling. This work deepens our understanding of the biological functions of RNA modifications in early development and provides a new perspective on RNA-mediated chromatin plasticity.
References:
[1] Xu, R., et al. (2022). Cell Stem Cell 29, 1051-1066.
[2] Liu, J., et al. (2021). Nature 591, 322-326.
[3] Chen, C., et al. (2021). Protein Cell 12, 455-474.
Original Article: https://doi.org/10.1016/j.stem.2025.10.003